API
How to access the INDYdb API?
Retrieving data from INDYdb is easy as using a "cURL GET request". Our database is structured using SQLite3, which is compatible with mutiple tools, such as: curl, wget, requests, jquery and many others.
Although there are several methods that can be used to do so, we highly recommend the use of a graphical interface that let's you build a query using visual cues. One such tool is Postman, which is freely available and will be used in this tutorial. There is both a web based version and local instance of Postman, but we highly recommend the use of the desktop version for improved stability.
So, without further ado, let's begin assembling our requests.
Image from https://www.postman.com/product/what-is-postman/
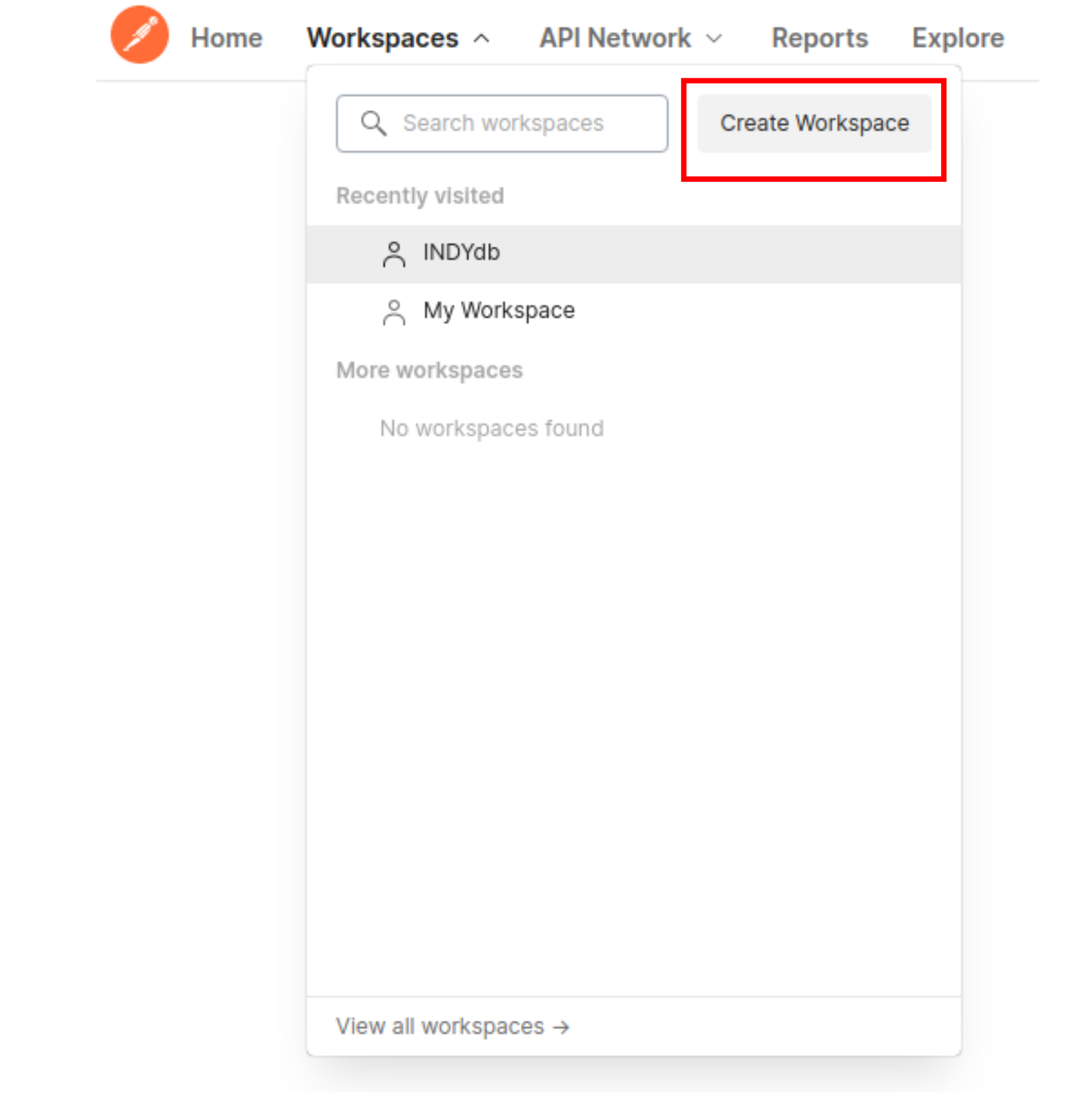
Create your new workspace
After creating your account, you'll find yourself in Postman home. Look for the the Workspaces tab and select "Create Workspace"
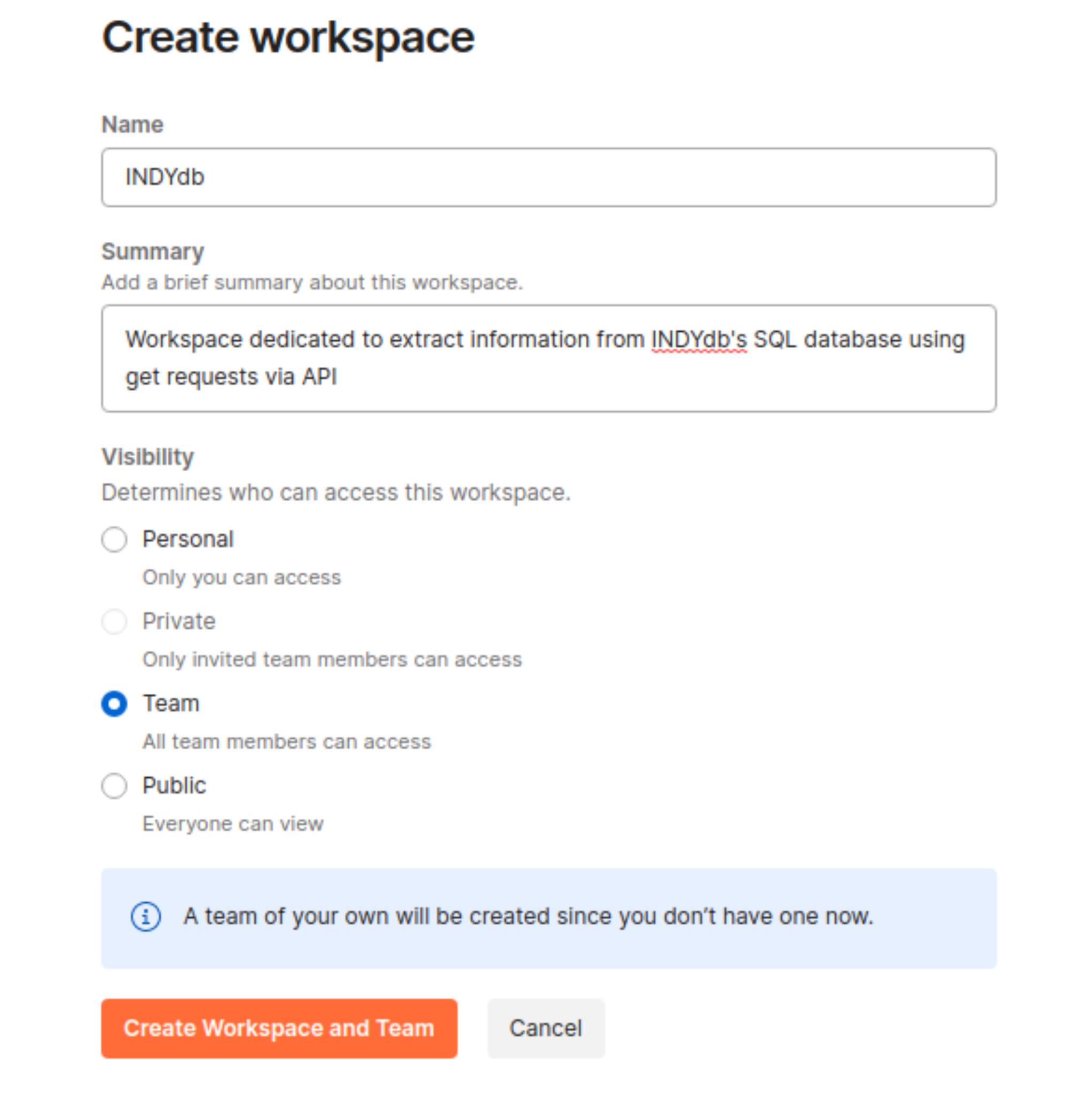
Fill in the information required
Fill the information required from creating you WP, such as name and a short summary. Don't forget to adjust the visibility options.
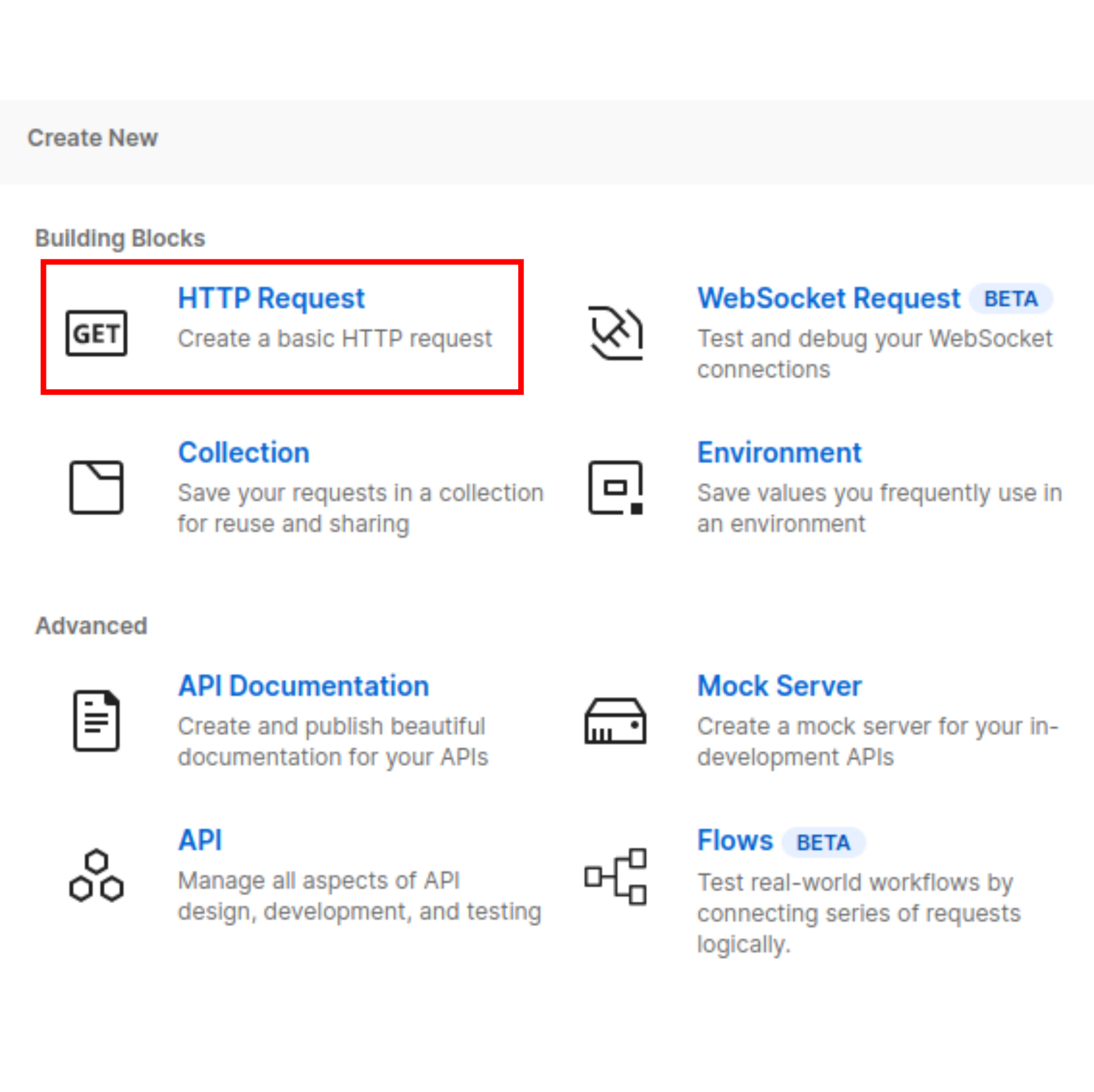
Create a new HTTP request
Once you load into your workspace, look for the "New" button on the top right corner of the screen and select HTTP request.
Generate your request
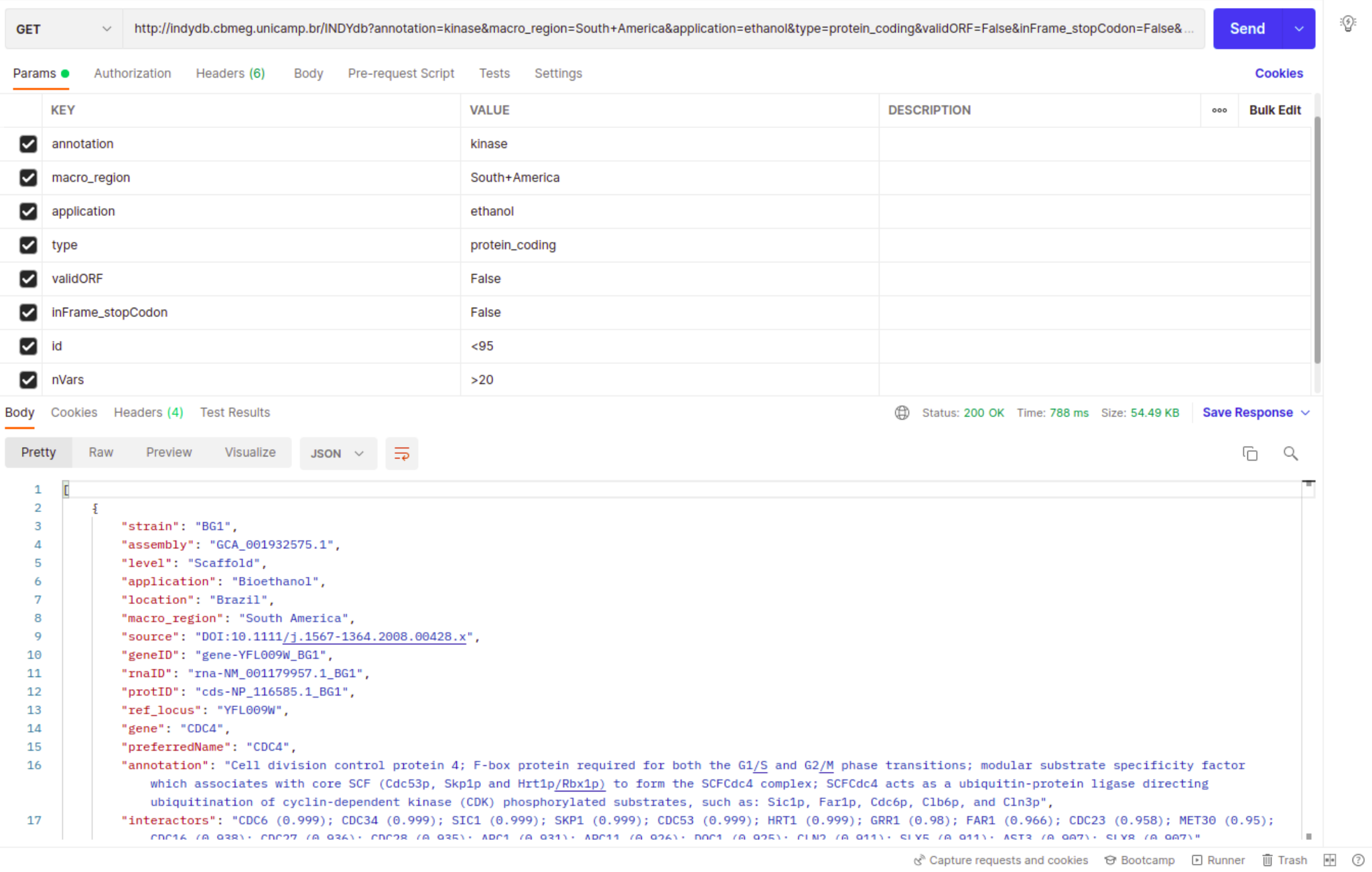
Once you open your HTTP request, retrieving the data is easy as filling a form. Just insert the INDYdb address with /INDYdb suffix added in the request URL with the GET option selected, fill in the key-value pairs and hit "Send". To get a list of all available query parameters, check out the API keys page.
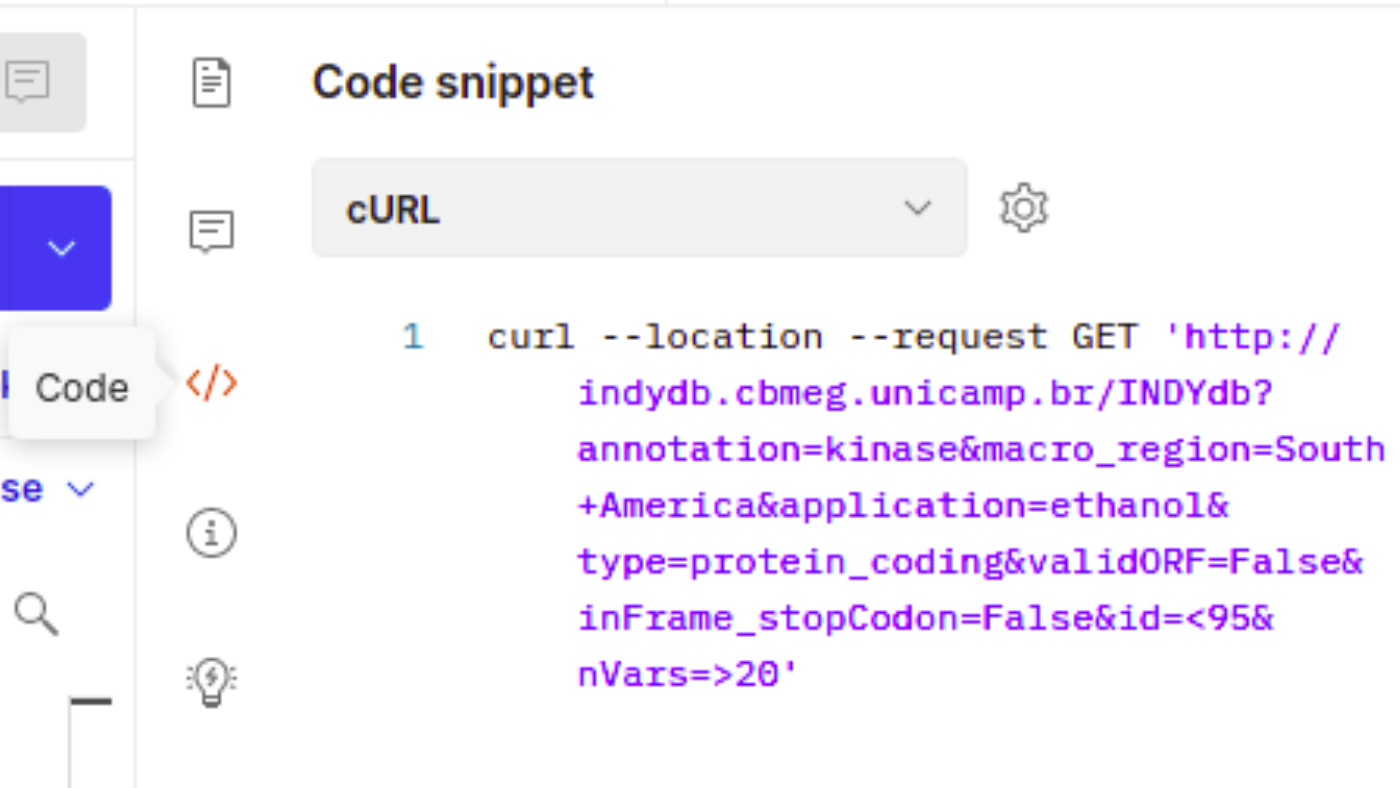
You can perform either a simple query with one parameter or perform complex queries using multiple parameter as shown in the example below, in which the query would read like: "Fetch all kinase-related genes from South American strains that produce ethanol, from that group return only the ones which the original gene is labeled as protein_coding but showed no valid ORF in the target sequence due to a out of frame stop codon. Further filter this group by selecting only the genes with less than 95% identity and at least 20 single nucleotide variants."
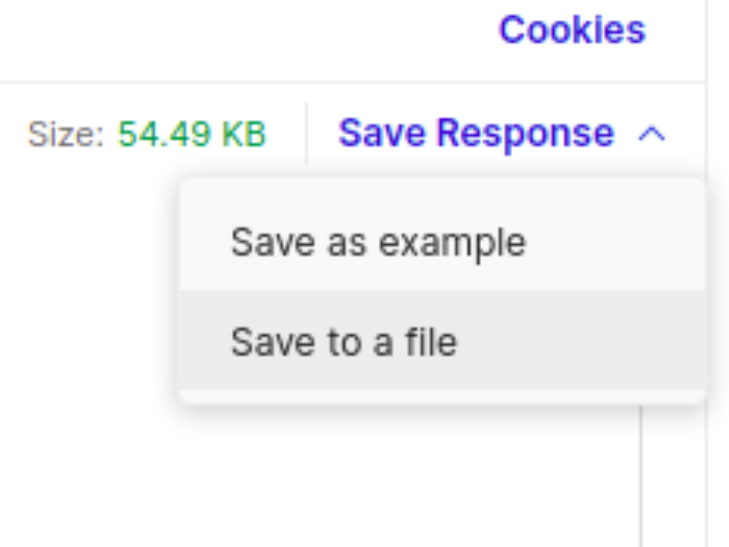
Save query to a local JSON file
After finishing your request, look for a button labeled as "Save Response" on the bar above the request results and select the "Save to a file" option. You'll be able to save your results as a JSON file, which can then be loaded into several data analysis platforms such as Python, R and Excel. However, we understand that not everyone is familiar with such data format, in this case we advise the use o file converter to shape the output into a CSV or XLSX format. There are several tools available for that, some are: csvconvert, csvjson, konklone's json to csv and konbert's json to csv.
Export query as a code snippet
Alternatively, you can also shape your query into a code that can be used in several other data analysis platforms. On the toolbar located on the right of the Postman app, you'll find a button called "Code". Opening this tab will allow you to read your get request as code snippets from multiple formats, from cURL and wget commands to more specific such as Python Requests or JavaScript jQuery, which can then be used as part of larger pipelines. As with the save to file option, this snippet will return JSON files that can be used in conjunction with data analyis tools or converted to other formats